Analysis modules
PDB Complex Interface Analysis
Identify interface residues in experimentally determined protein complexes directly from the PDB.
User Complex Interface Analysis
Analyze protein–protein interfaces in your own structural models by uploading a custom complex.
Patch Analysis
Predict interacting residues on monomeric protein surfaces using patch scoring methods.
Cluster Analysis
Group predicted interacting residues into functional clusters on the protein surface.
Surface visualization
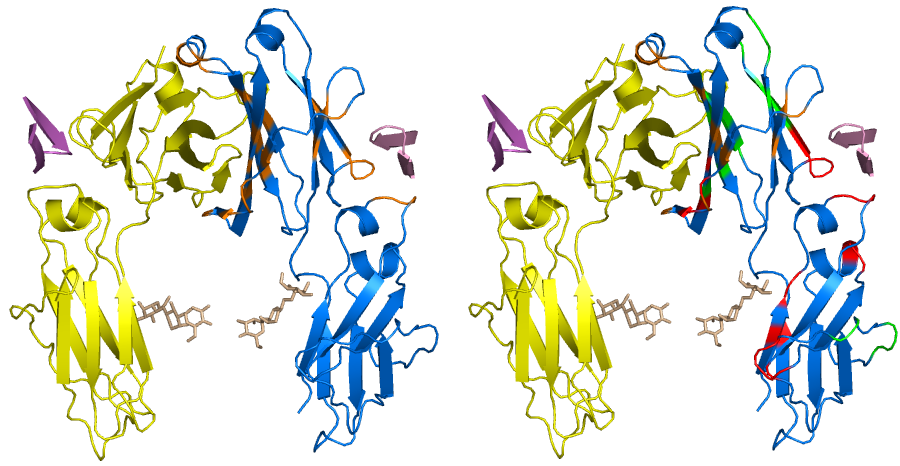
References
- Negi S.S., Schein C.H., Oezguen N., Power T.D., Braun W. InterProSurf: a web server for predicting interacting sites on protein surfaces. Bioinformatics. 2007;23:3397–3399.
- Negi S.S., Braun W. Statistical analysis of physical–chemical properties and prediction of protein–protein interfaces. J Mol Model. 2007;13:1157–1167.
- Negi S.S., Kolokoltsov A.A., Schein C.H., Davey R.A., Braun W. Determining functionally important amino acid residues of the E1 protein of Venezuelan equine encephalitis virus. J Mol Model. 2006;12:921–929.
Disclaimer
This service is provided freely to the scientific community for research purposes. There is no warranty, expressed or implied. The user assumes all responsibility for the use, misuse, or inability to use this service. The Sealy Center for Structural Biology is not liable for any damages arising from its use.